- Location:
-
1 H2.1; 1 70.53 cM
- Exon count:
- 6
| Annotation release |
Status |
Assembly |
Chr
|
Location |
| RS_2024_02 |
current |
GRCm39 (GCF_000001635.27) |
1 |
NC_000067.7 (163072688..163142714, complement) |
| 108.20200622 |
previous assembly |
GRCm38.p6 (GCF_000001635.26) |
1 |
NC_000067.6 (163245119..163315145, complement) |
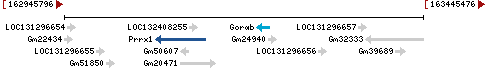
Chromosome 1 - NC_000067.7